|
9/19/2023 0 Comments Cytoscape 2.8 Our results on 12 signaling pathways from the NetPath database demonstrate that the application can be used for indicating proteins which may play significant roles in a pathway by finding the strongest path(s) in the PPI or signaling network. This application can be used on the built-in human and mouse PPI or signaling databases, or any user-provided database for some organism. The application can also identify any activating or inhibitory regulatory paths between two distinct sets of transcription factors and target genes. Given a set of proteins, StrongestPath can extract a set of possible interactions between the input proteins, and expand the network by adding new proteins that have the most interactions with highest total confidence to the current network of proteins. When there are different levels of confidence over the interactions, the application is able to process them and identify the cascade of interactions with the highest total confidence score. Save a network to your GenomeSpace cloud storage.Ĭlicking File>GenomeSpace provides several options for managing your files in your GenomeSpace cloud storage and using GenomeSpace to launch other applications.ĭelete a file from your GenomeSpace cloud storage.ĭownload a file in your GenomeSpace cloud storage to your local machine.StrongestPath is a Cytoscape 3 application that enables the analysis of interactions between two proteins or groups of proteins in a collection of protein–protein interaction (PPI) network or signaling network databases. Saves a session file (CYS), containing all networks, attributes (for node/edge/network), Desktop states (selected/hidden nodes and edges, window sizes), Properties, some plugin states, and Visual Styles in your present working state to your GenomeSpace cloud storage. Uploads a file from your local machine to the root level of your GenomeSpace cloud storage. Annotations in Cytoscape are stored as a set of ontologies (a set of controlled vocabulary terms that annotate the objects).The standard file formats used in Cytoscape are OBO and Gene Association.Ĭlicking File>Export>GenomeSpace provides options for exporting files to GenomeSpace. Loads an annotation/ontology file from your GenomeSpace cloud storage. A session file (CYS) contains all networks, attributes (for node/edge/network), Desktop states (selected/hidden nodes and edges, window sizes), Properties, some plugin states, and Visual Styles present in a given working state. Loads a CYS file from your GenomeSpace cloud storage. Starts the import from your GenomeSpace cloud storage of delimited text or single-sheet MS Excel workbook containing attribute data tables that are not formatted into Cytoscape node or edge attribute file formats. The load dialog allows you to opt to either Load node attributes or Load edge attributes. Loads a file containing attributes for a CyNetwork from your GenomeSpace cloud storage. Loads attributes (for node/edge/network) from your GenomeSpace cloud storage. Starts the import from your GenomeSpace cloud storage of a network from a delimited text file or single-sheet MS Excel workbook. Loads a network from your GenomeSpace cloud storage. You can also Reset Password on your existing GenomeSpace account.Ĭlicking File>Import>GenomeSpace provides several options for importing files from GenomeSpace. If you start from your GenomeSpace-enabled JNLP of Cytoscape and want to log into GenomeSpace in order to access your Amazon cloud storage or other GenomeSpace tools, you will be asked to log in with your GenomeSpace username and password. 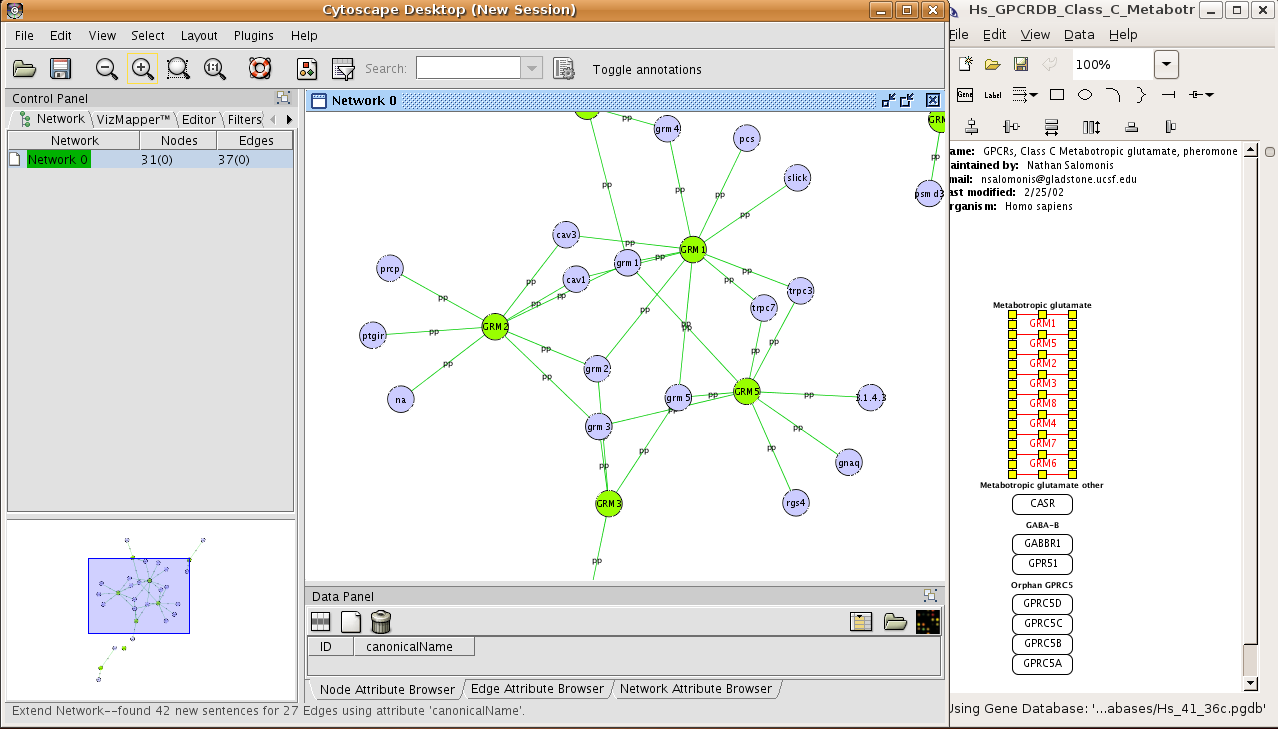
When you launch Cytoscape from GenomeSpace, your login should be handled seamlessly. You can download plug-ins for your Cytoscape by choosing Plugins>Manage Plugins.Ĭytoscape has several GenomeSpace menu items. Like several other applications in the GenomeSpace suite of tools, Cytoscape can be expanded with plug-ins. If you launch Cytoscape from the Cytoscape website, at this time it will not be the version that is linked to GenomeSpace. 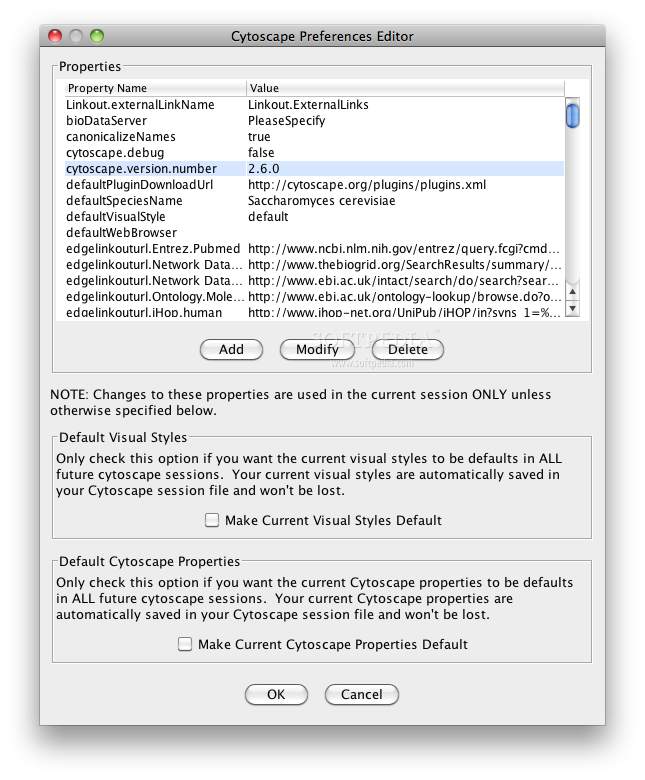
Once it's downloaded to your machine (after a launch from GenomeSpace), you can start it locally (by opening the JNLP) or launch from GenomeSpace. Additional features are available as plugins.Ĭytoscape is a Java-based application that downloads to your local machine when you launch it from GenomeSpace. Cytoscape core distribution provides a basic set of features for data integration and visualization. In the meantime, below is the help from the previous revision (Cytoscape 2.8)Ĭytoscape is an open-source bioinformatics software platform for visualizing molecular interaction networks and biological pathways, and integrating these networks with annotations, gene expression profiles, and other state data. New Documentation for Cytoscape 3 is on its way. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
